HEPData Output Formats
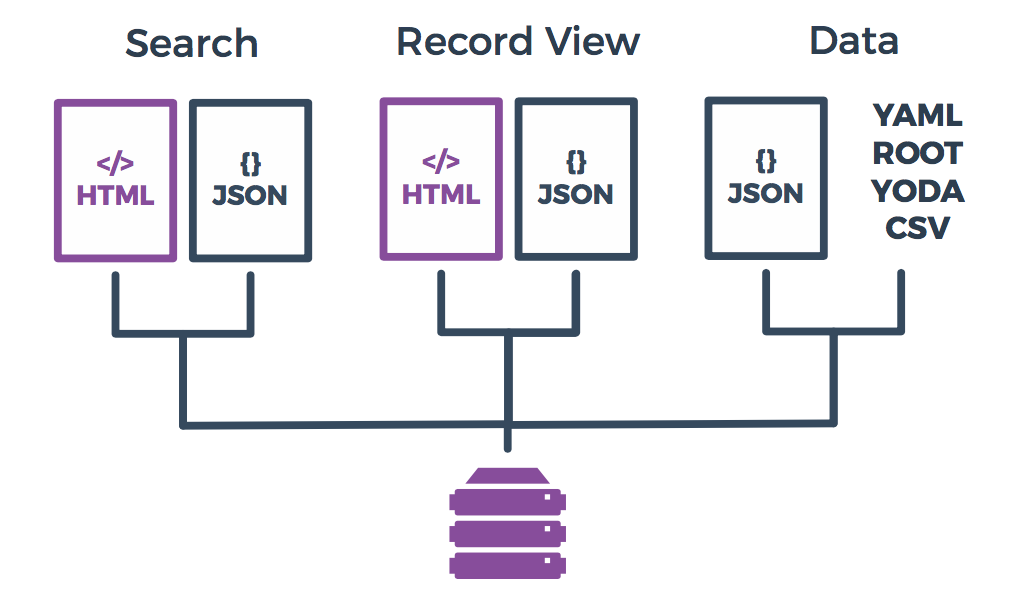
In addition to browsing HEPData, there are various ways to interact with it programatically and to retrieve data in different formats:
HEPData Output Formats

In addition to browsing HEPData, there are various ways to interact with it programatically and to retrieve data in different formats:
JSON Endpoints
Aside from the normal HTML view of HEPData, most of our pages and data have a JSON equivalent. The JSON format allows programmatic access, from scripts in different languages such as Python or from applications such as Mathematica.
Simply add &format=json to the search URL or detailed record URL to get the results back in JSON.
If a user goes to the search page and filters on Collaboration: ATLAS, they will land at the URL:
By default, the search results are returned with up to 10 records on each page (size=10), but this number can be
altered by passing size=1 up to size=100 in the search URL.
Multiple URL arguments should each be separated by a & symbol.
Record pages also have a JSON view, for example:
The light view reduces the size of the JSON by removing data tables and is useful for getting information about the whole record.
Data file formats
Conversion to various export formats is provided by a separate extensible package, hepdata-converter. Current output formats are:
hepdata_doi and table_doi fields.
.root file with each table in a
separate directory. For numeric data, a TGraphAsymmErrors object is written for each
dependent variable. If the data has finite bin widths, then also separate TH1F
objects are written for the central value of the data points and each of the uncertainties. If
there is more than one independent variable, the appropriate
ROOT object is chosen, for example,
a TH2F or TH3F instead of a TH1F object.
yoda output is now
in the YODA2 format, but there is still an option yoda1 to output the legacy YODA1 format.
Alternatively, the output can be given in the yoda.h5 format, designed for parallel computing.
To download in a specific format, append a format query parameter to the URL of a
record, using one of:
?format=json?format=original?format=yaml?format=csv?format=root?format=yoda?format=yoda1?format=yoda.h5Optional parameters can also be added (separated by a & symbol):
table=Table%201: provide the table name in order to download a specific table instead of all tables.
Special characters should generally be URL-encoded,
for example, %20 for a space. Omitting spaces also works, for example, table=Table1.
version=1: specify a particular version of a record. If omitted, the latest version
will be returned.
light=true: when using format=json for a whole submission, this omits
the data tables from the response.
rivet=ALICE_2016_I1419244: when using format=yoda, format=yoda1, or format=yoda.h5, specify the desired
Rivet analysis name to be written in the YODA files if it does not match the automatically generated name.
qualifiers=true: when using format=yoda, format=yoda1, or format=yoda.h5, will include metadata on
the table qualifiers in the exported YODA files as annotations, for potential use cases outside of Rivet analyses.
For example, https://www.hepdata.net/record/ins1419244?format=yaml&table=Table1&version=1 returns Table 1 of version 1 of the record, in YAML format.
Content Negotiation
HEPData record pages support content negotiation to allow automated downloads of JSON-LD metadata and of resource files attached to a submission.
HEPData record pages include JSON-LD metadata to provide machine-readable information about the record.
To download just the JSON-LD for a record page, use the HTTP header
Accept: application/ld+json in your request, e.g.:
curl -OJLH "Accept: application/ld+json" https://www.hepdata.net/record/ins1419244
If you are using the DOI for a record rather than the HEPData URL, passing
Accept: application/ld+json will give you the JSON-LD metadata directly from DataCite, which is
less complete than the metadata produced by HEPData. To get the HEPData metadata
via the DOI, use the header Accept: application/vnd.hepdata.ld+json, e.g.:
curl -OJLH "Accept: application/vnd.hepdata.ld+json" https://doi.org/10.17182/hepdata.72886.v2
DOIs for resource files relating to a HEPData submission direct users to a landing page such as
https://www.hepdata.net/record/resource/1935437?landing_page=true.
Omitting the landing_page=true URL argument will return JSON metadata for the resource file,
while replacing it with view=true will download the resource file.
To download a resource file directly from its DOI, pass the relevant
media type
in the Accept header. For example, for DOI 10.17182/hepdata.89408.v3/r2:
curl -OJLH "Accept: application/x-tar" https://doi.org/10.17182/hepdata.89408.v3/r2
If you pass an Accept header that is not valid for the given resource, you will receive a
response with a 406 Not Acceptable
status code with an error message in JSON format such as:
{
"file_mimetype": "application/x-tar",
"msg": "Accept header value 'application/zip' does not contain a valid media type for this resource. Expected Accept header to include one of 'application/x-tar', 'text/html', 'application/ld+json', 'application/vnd.hepdata.ld+json'"
}
The file_mimetype field provides the correct media type to use in the Accept
header to download the resource.
Note that both the curl command and the Python requests module have a
default Accept header of */*, different from
most web browsers
where */* has a lower weighting than text/html. The HEPData code will return the content
directly if */* has weight 1 in a request to a landing page, therefore it is not strictly necessary
to specify an explicit Accept header when using curl or Python requests,
although it is still recommended to do so.
Some text from this page is adapted from:
HEPData: a repository for high energy physics data
Eamonn Maguire (CERN), Lukas Heinrich (New York U.), Graeme Watt (Durham U., IPPP)
e-Print: 1704.05473 [hep-ex]
DOI: 10.1088/1742-6596/898/10/102006
Published in: J.Phys.Conf.Ser. 898 (2017) 10, 102006